
Researchers from Imperial College London, CABI and the University of Oxford have sequenced the genome of Alexander Fleming’s original Penicillium strain using samples that were frozen alive more than fifty years ago.
Dr Matthew Ryan, Curator of the Genetic Resource Collection at CABI, shared his expertise and samples of Fleming’s original Penicillium strain with lead researcher Professor Timothy Barraclough, from the Department of Life Sciences at Imperial and the Department of Zoology at Oxford, to enable the work to take place.
Alexander Fleming famously discovered the first antibiotic, penicillin, in 1928 while working at St Mary’s Hospital Medical School, which is now part of Imperial College London. The antibiotic was produced by a mould in the genus Penicillium that accidentally started growing in a Petri dish.
One strain from Fleming’s investigations was transferred to the Imperial Mycological Institute, IMI (part of CABI) in 1945. Dr Ryan said the strain Fleming originally sent to Charles Thom was transferred to CABI in 1950 and the first strain to be used to produce pure cultures of penicillin by ‘submerged culture production’ was from Belgium (NRRL 832), deposited at CABI in 1947.
The team used the new genome to compare Fleming’s mould with two strains of Penicillium from the US that are used to produce the antibiotic on an industrial scale. The results, published today in Scientific Reports, reveal that the UK and US strains use slightly different methods to produce penicillin, potentially suggesting new routes for industrial production.
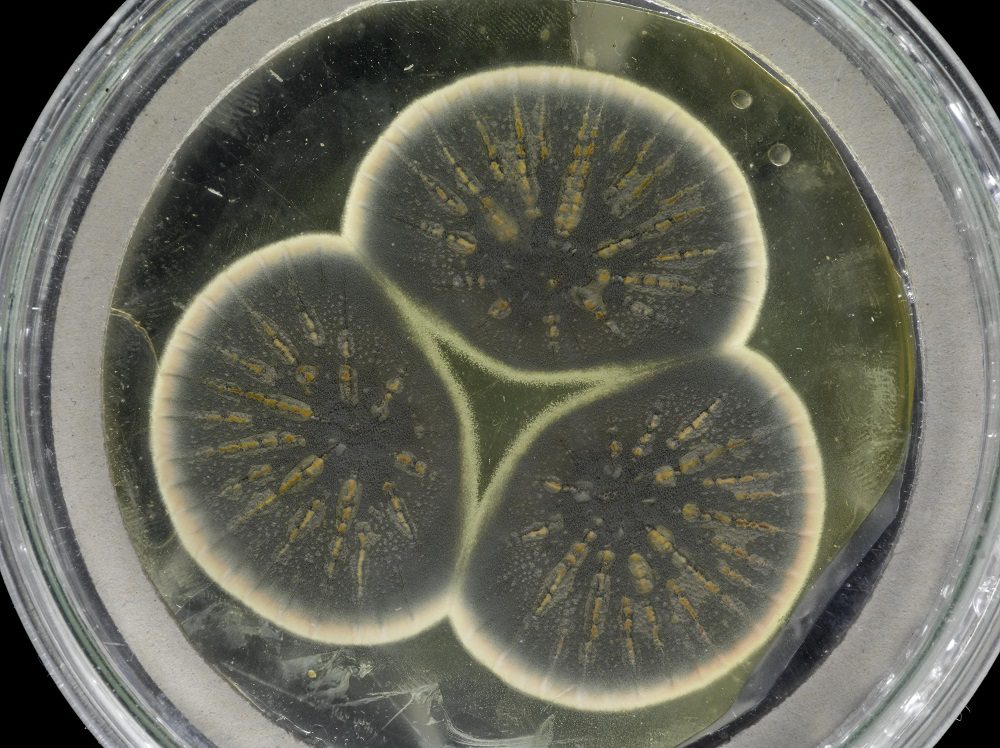
Researchers have sequenced the genome of Fleming’s original Penicillium strain using samples that were frozen alive more than fifty years ago (Credit: CABI).
Professor Barraclough said: “We originally set out to use Alexander Fleming’s fungus for some different experiments, but we realised, to our surprise, that no-one had sequenced the genome of this original Penicillium, despite its historical significance to the field.”
Although Fleming’s mould is famous as the original source of penicillin, industrial production quickly moved to using fungus from mouldy cantaloupes in the US. From these natural beginnings, the Penicillium samples were artificially selected for strains that produce higher volumes of penicillin.
The team re-grew Fleming’s original Penicillium from a frozen sample kept at the culture collection at CABI and extracted the DNA for sequencing. The resulting genome was compared to the previously published genomes of two industrial strains of Penicillium used later in the US.
The researchers looked in particular at two kinds of genes: those encoding the enzymes that the fungus uses to produce penicillin; and those that regulate the enzymes, for example by controlling how many enzymes are made.
In both the UK and US strains, the regulatory genes had the same genetic code, but the US strains had more copies of the regulatory genes, helping those strains produce more penicillin.
However, the genes coding for penicillin-producing enzymes differed between the strains isolated in the UK and US. The researchers say this shows that wild Penicillium in the UK and US evolved naturally to produce slightly different versions of these enzymes.
Moulds like Penicillium produce antibiotics to fight off microbes, and are in a constant arms race as microbes evolve ways to evade these defences. The UK and US strains likely evolved differently to adapt to their local microbes.
Microbial evolution is a big problem today, as many are becoming resistant to our antibiotics. Although the researchers say they don’t yet know the consequences of the different enzyme sequences in the UK and US strains for the eventual antibiotic, they say it does raise the intriguing prospect of new ways to modify penicillin production.
First author Ayush Pathak, from the Department of Life Sciences at Imperial, said: “Our research could help inspire novel solutions to combatting antibiotic resistance. Industrial production of penicillin concentrated on the amount produced, and the steps used to artificially improve production led to changes in numbers of genes.
“But it is possible that industrial methods might have missed some solutions for optimising penicillin design, and we can learn from natural responses to the evolution of antibiotic resistance.”
Additional information
Main image: The freeze-dried Fleming strain from which the Penicillium fungus was grown and genome sequenced (Credit: CABI).
Full paper reference
Ayush Pathak, Reuben W. Nowell, Christopher G. Wilson, Matthew J. Ryan and Timothy G. Barraclough, ‘Comparative genomics of Alexander Fleming’s original Penicillium isolate (IMI 15378) reveals sequence divergence of penicillin synthesis genes,’ 24 September 2020, Scientific Reports, DOI: 10.1038/s41598-020-72584-5
The paper can be read pen access here: https://www.nature.com/articles/s41598-020-72584-5
This story was written from an original press release from Imperial College London which can be read in full here: draft here: https://www.imperial.ac.uk/news/204713/genome-alexander-flemings-original-penicillinproducing-mould/
Science Museum Blog
Dr Matthew Ryan explores Fleming’s Penicillium and the potential of microorganisms in the blog ‘Keeping History Alive!’ published by the Science Museum. Read more here: https://blog.sciencemuseum.org.uk/keeping-history-alive/
Related News & Blogs




